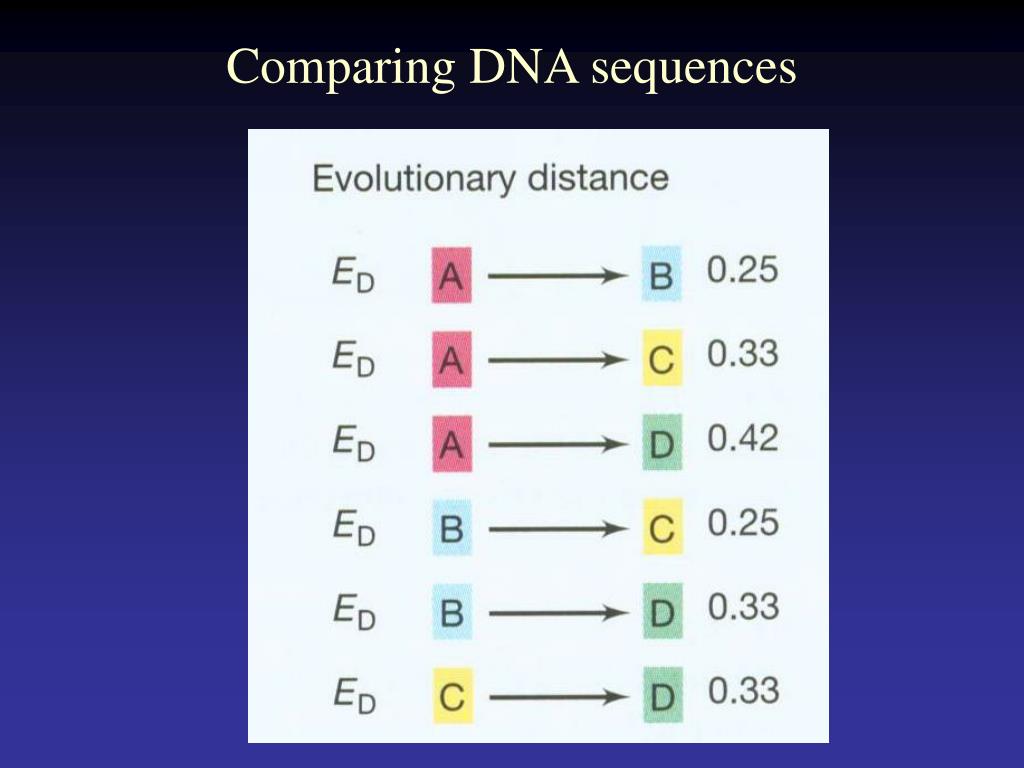
 
These reads are compared to each other and those sharing the same DNA sequence are grouped together.Assembly of a de novo sequence begins with a large number of short sections or “reads” of DNA.Illustration showing the difference between single and paired-end reads. Paired-end reads are particularly useful when assembling a de novo sequence as they provide long-range information that you wouldn’t otherwise have in the absence of a gene map. This makes it easier to assemble them into a continuous DNA sequence. The key advantage of paired-end reads is that scientists know how far apart the two ends are. The distance between paired-end reads can be anywhere between 200 base pairs and several thousand. Paired-end reads are where both ends of a fragment of DNA are sequenced.These sequences can then be joined together by finding overlapping regions in the sequence to create the full DNA sequence. Single reads are where one end or the whole of a fragment of DNA is sequenced.Producing a gene map can be an expensive process, so some assembly programmes rely on data consisting of a mix of single and paired-end reads (see illustration below):.To help assemble a de novo sequence a physical gene map can be developed before sequencing to highlight the “landmarks” so the scientists know where sections of DNA are located in relation to each other.If you know that the new species is very similar to another species that does have a reference genome, it is possible to assemble the sequence using a similar genome as a guide. 
In de novo assembly there is no existing reference genome sequence for that species to use as a template for the assembly of its genome sequence.De novo sequencing is when the genome of an organism is sequenced for the first time.This is primarily carried out for de novo sequences. It is an attempt to reconstruct the original genome. Assembly involves taking a large number of DNA reads, looking for areas in which they overlap with each other and then gradually piecing together the ‘jigsaw’.Alignment is when the new DNA sequence is compared to existing DNA sequences to find any similarities or discrepancies between them and then arranged to show these features.This is done using processes called alignment and assembly:.Put the pieces together in the correct order to construct the complete genome sequence and identify any areas of interest.Like the pieces of a jigsaw puzzle, these DNA reads are jumbled up so we need to piece them together and put them in the correct order to assemble the genome sequence.To put this into perspective, once a human genome has been fully sequenced we have around 100 gigabases (100,000,000,000 bases) of sequence data.High coverage means that after sequencing DNA we have lots and lots of pieces of DNA sequence (reads).It is also much cheaper to carry out sequencing to a higher coverage than it was at the time of the Human Genome Project.Having a higher coverage reduces the likelihood of there being gaps in the final assembled sequence. Some sequencing technologies deal with shorter reads of DNA which means that gaps are more likely to occur when the genome is assembled.Although most current sequencing techniques are now faster than they were during the Human Genome Project, some sequencing technologies have a higher error rate.Coverage has increased because of a few reasons: During the Human Genome Project coverage was only between 5- and 10-fold and used a different sequencing technology to those used today.30- to 50-fold coverage is currently the standard used when sequencing human genomes to a high level of accuracy.Effectively, the more times you sequence, or “read”, the same section of DNA, the more confidence you have that the final sequence is correct.For example, 30 times (30-fold) coverage means each base is sequenced 30 times. So, to account for the errors that could potentially occur, each base in the genome is sequenced a number of times over, this is called coverage.The technology of DNA sequencing is not 100 per cent accurate and therefore there are likely to be errors in the DNA sequence that is produced. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
